Yu S. Huang

Shanghai/Beijing/Wuxi, China
Ph.D. with 15+ years of leadership in computational biology, AI drug discovery, genomic/protein AI modeling, and high-performance computing (HPC) infrastructure. Expert in generative AI, protein language models, multimodal fusion, structure-based drug design. Proven track record in defining technical strategy, building end-to-end AI platforms from scratch, leading cross-functional teams, managing academic‑industry collaborations, and driving publications & IP strategy. Led the development and industrial deployment of AI models for cancer early detection, virtual screening, and molecular generation.
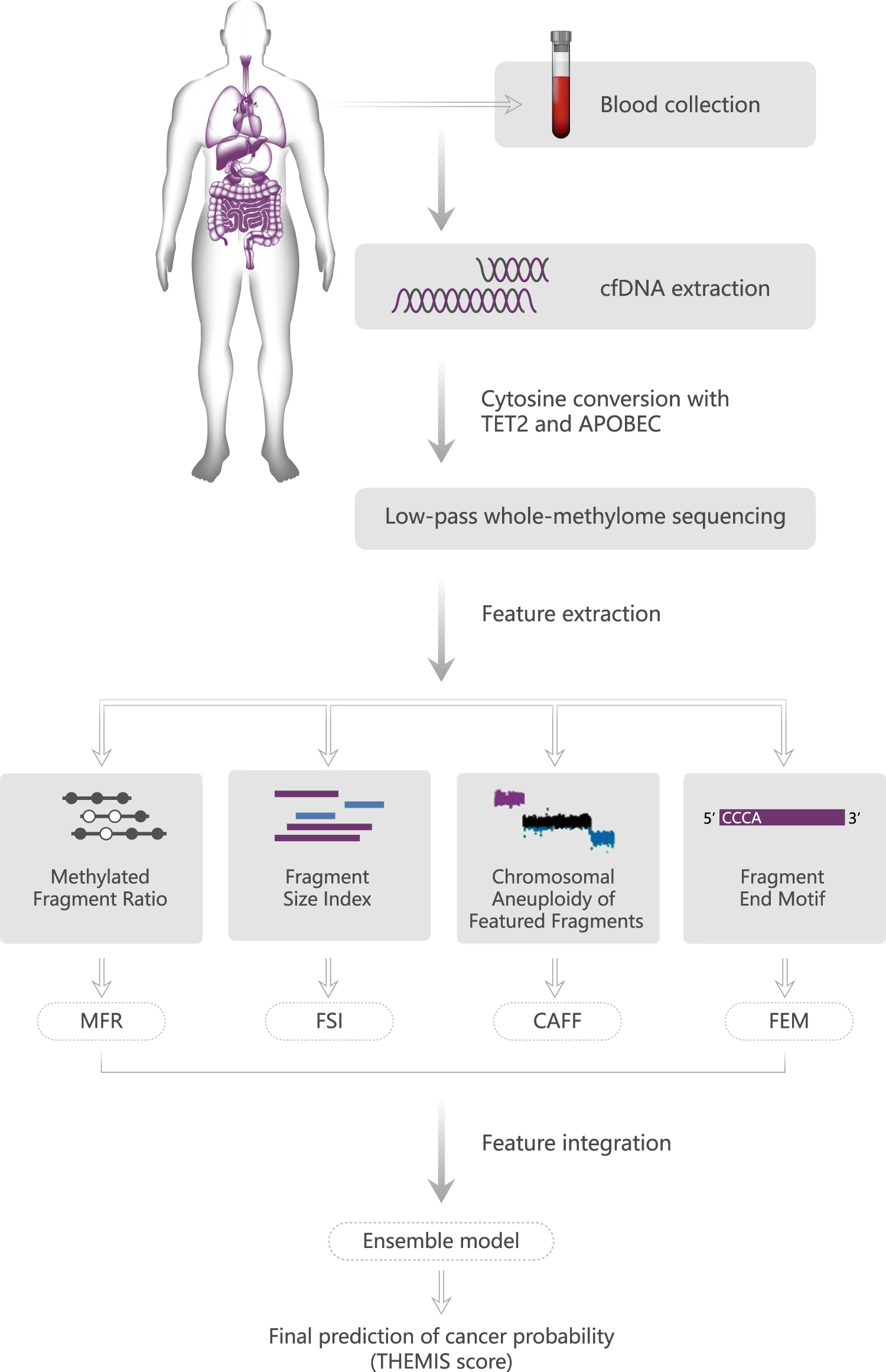
Since 2022, I have served as Senior Director 资深总监计算生物学 at 臻和 Genecast Corp Ltd.. My main responsibility is to define and execute long-term technical strategy for AI-driven precision oncology, aligned with corporate product pipelines and business goals. Lead the development of multimodal AI platforms integrating sequence, structure, and epigenomic data for non-invasive cancer detection. Built enterprise-grade AI computing infrastructure (K8s, PyTorch, distributed storage, high-speed interconnect) to support large-scale computing. Lead and mentor a high-performance team of algorithm scientists, bioinformaticians, and software engineers to deliver end-to-end solutions from in silico modeling to experimental validation. Led cross-disciplinary team management and promoted tight integration between computational models and experimental biology. External scientific engagement, conference presentations, high-impact publications, and IP strategy; drove research-to-product translation.
From 2015 to 2021, I was Professor and Principal Investigator at Drug Discovery and Design Center, Shanghai Institute of Materia Medica, Chinese Academy of Sciences (中科院). I led the establishment of AI-driven computational biology and drug discovery center and built a mature structure-based drug design & virtual screening system. Developed Fergie (VAE-based small molecule generation) and Deffini (structure-based virtual screening DNN) to enable structure-guided drug design at scale, with one kinase inhibitor molecule entering the PCC phase. Developed core algorithms for genomic variant calling, copy number analysis, and methylation sequencing to support early-stage innovative drug R&D. Directed national/provincial research projects, built academic-industry partnerships, and delivered high-impact publications. During this time, I also taught courses on Artificial Intelligence and Machine Learning, Julia (a scientific computing language), Matrix Computations, Optimization.
My career also includes experience as a Bioinformatics Scientist at Illumina Inc. (San Diego) and a Postdoctoral Researcher in Human Genetics at UCLA. I earned my Ph.D. in Computational Biology and Bioinformatics from USC, specializing in machine learning, statistical learning, graph theory, population genetics. Inside a Ph.D. program founded by mathematician and founding father of computational biology M.S. Waterman, I gained a comprehensive understanding of machine learning, statistics, probability, stochastic processes.
I received my B.S. in Biology from Fudan University in July 2003, during which I became proficient in C/C++, PostgreSQL, Java, Python, and Linux systems. My fascination with computers began with writing my first BASIC program on an Intel-8088 PC in the 8th grade.
In my spare time, I enjoy reading, surfing, skateboarding, and snowboarding.
- GitHub 代码库: github.com/polyactis
- ORCID 0000-0001-5967-4948, ResearchGate
- 小红书: polyactis
- Expertise: Machine/Deep/Statistical Learning, Bioinformatics, AI Drug Design, Optimization, Distributed Computing


latest posts
| Aug 06, 2025 | AI赋能肿瘤基因诊断!黄宇博士深度解析:技术创新如何破解临床痛点 - 中国抗癌协会TBM医研企座谈会 |
|---|---|
| Jun 12, 2025 | Bayesian p-value/贝叶斯p-value |
| Sep 02, 2024 | How to calculate the derivative of a matrix multiplication such as WHW |
| Apr 18, 2024 | 贝叶斯统计 Statistical Rethinking v2 in Julia 讲解视频 |
| Mar 15, 2024 | 7 Julia correctness issues are now fixed. |